What is BUGSS?
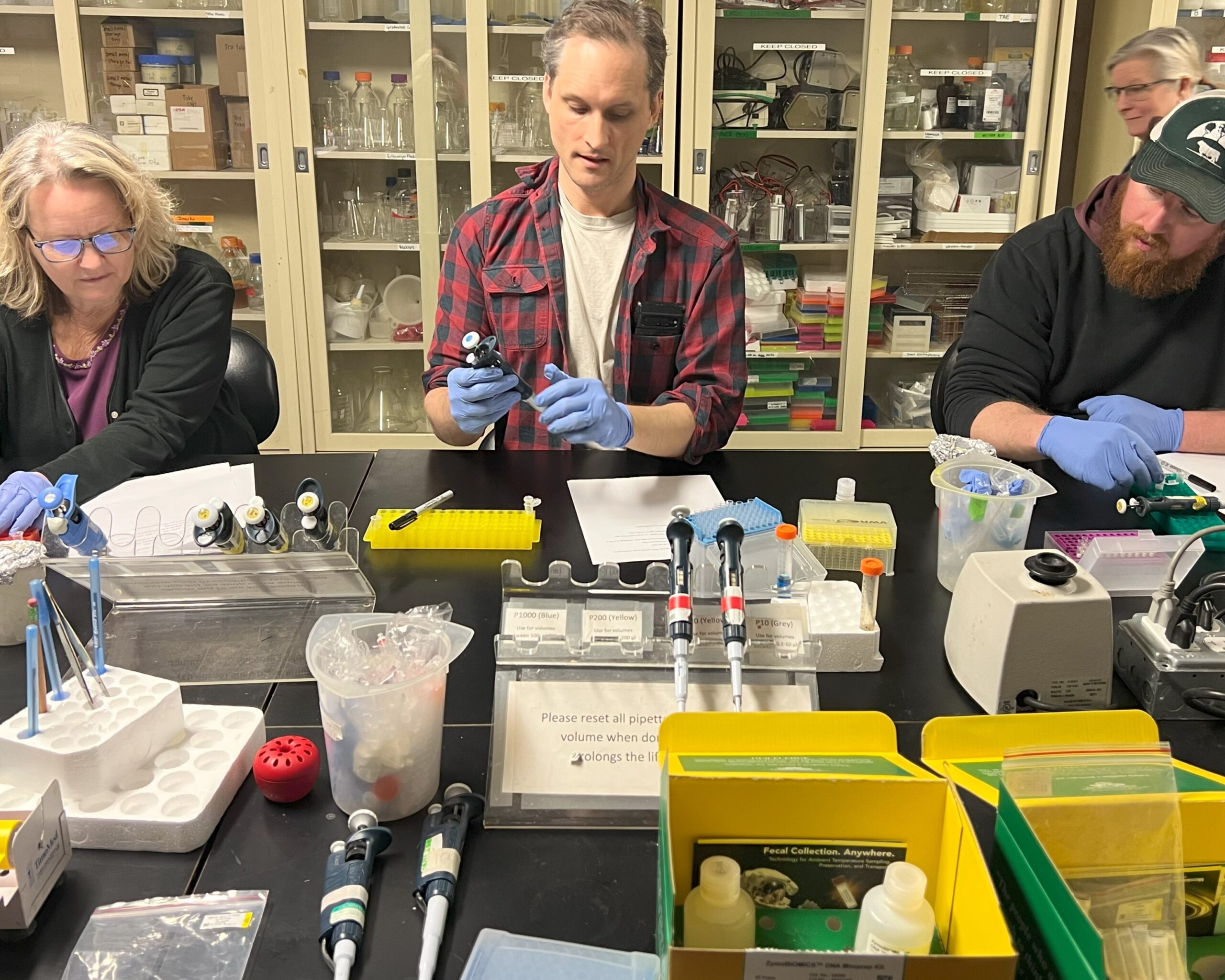
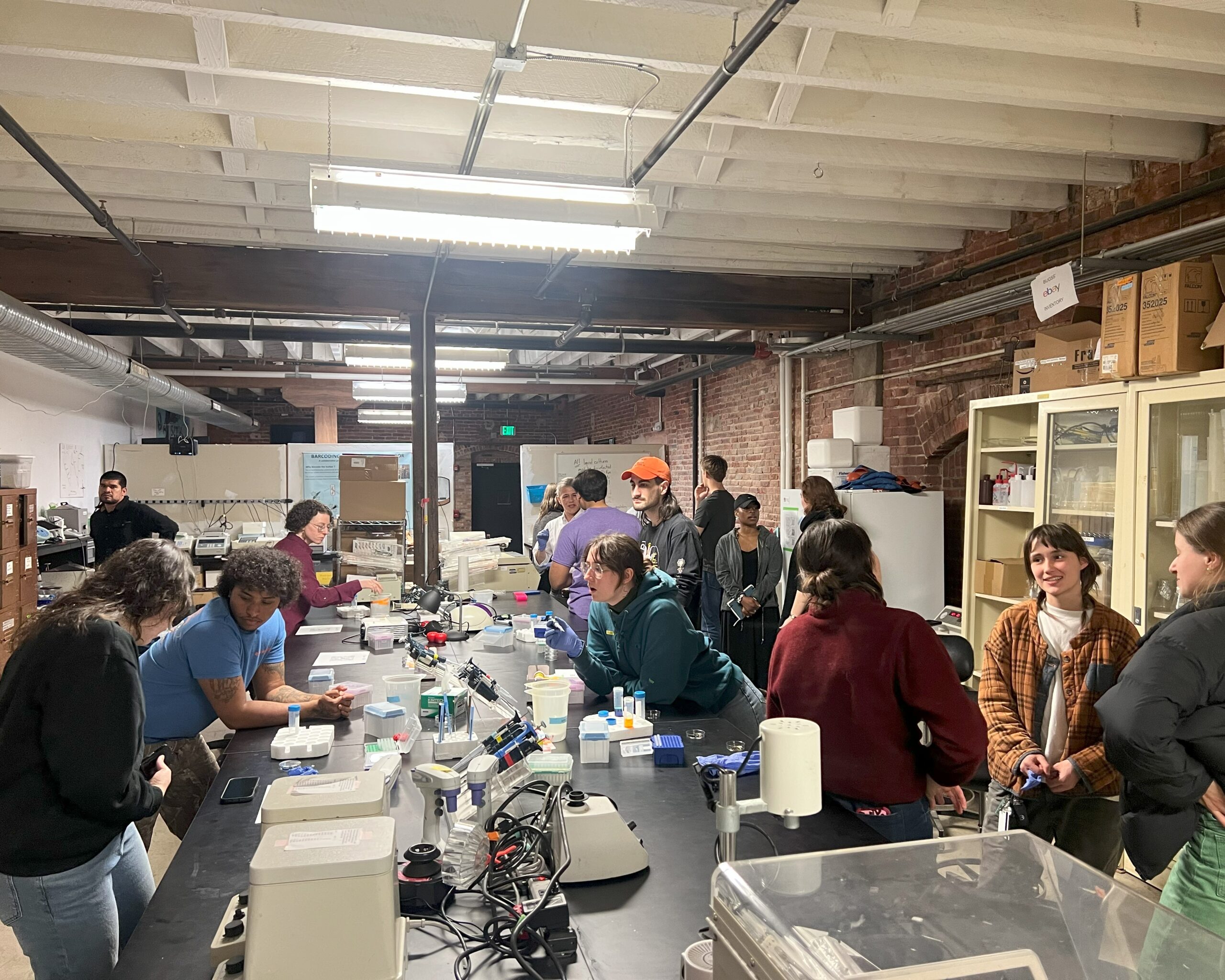
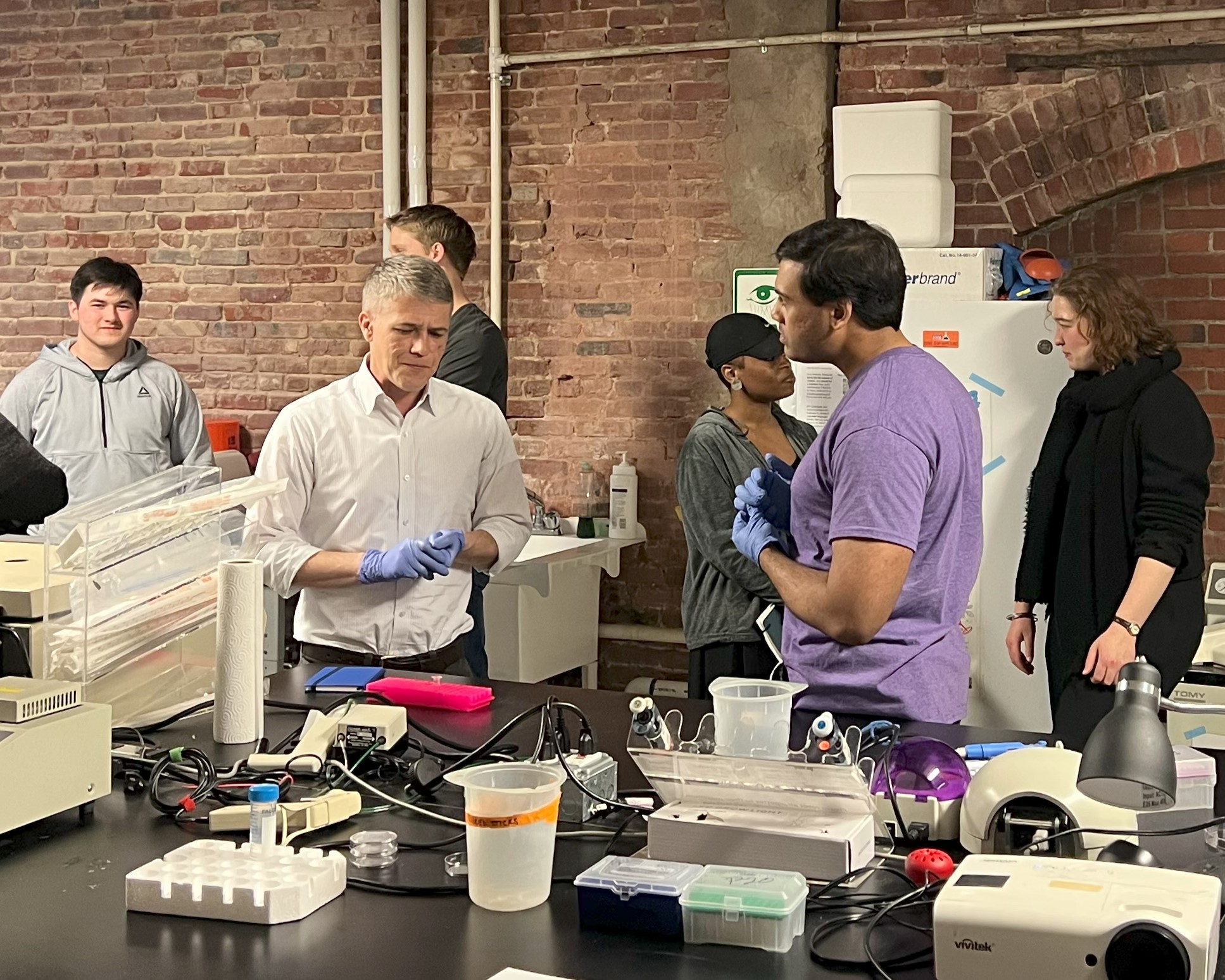
BUGSS is a non-profit public laboratory offering classes, seminars, and lab access so that anyone can safely and affordably investigate the living world. By democratizing these technologies we hope to facilitate more nuanced dialogue and exploration of the incredible potential, as well as limitations and ethical issues.
Upcoming Events
These are the new and exciting events coming up soon! Learn more about each one on our Eventbrite pages!